PROPTIMUS
LIVE
Constrained α-carbons optimisation
of protein structures
About PROPTIMUS LIVE
PROPTIMUS LIVE is a freely available application for the local constrained α-carbons optimisation of protein structures.
It is powered by the GFN-Force-Field, accelerated by a divide-and-conquer RAPHAN approach.
How PROPTIMUS LIVE works
Repairs non-physically predicted atoms with the pdb2pqr tool,
which may be present in ML-predicted protein structures.
Adds hydrogens for user-specified pH using the pdb2pqr tool
if the input structure does not contain them.
Optimises protein structure with constrained α-carbons
using the GFN-Force-Field.
Why use PROPTIMUS LIVE
High-quality results because of the use of an almost
quantum-mechanics accurate GFN-Force-Field.
Fast calculations within minutes due to the acceleration
by divide-and-conquer RAPHAN approach.
Free and open to all users—no login required.
How PROPTIMUS LIVE improves structures?
Original structures are grey, while those optimized by PROPTIMUS LIVE are colored.
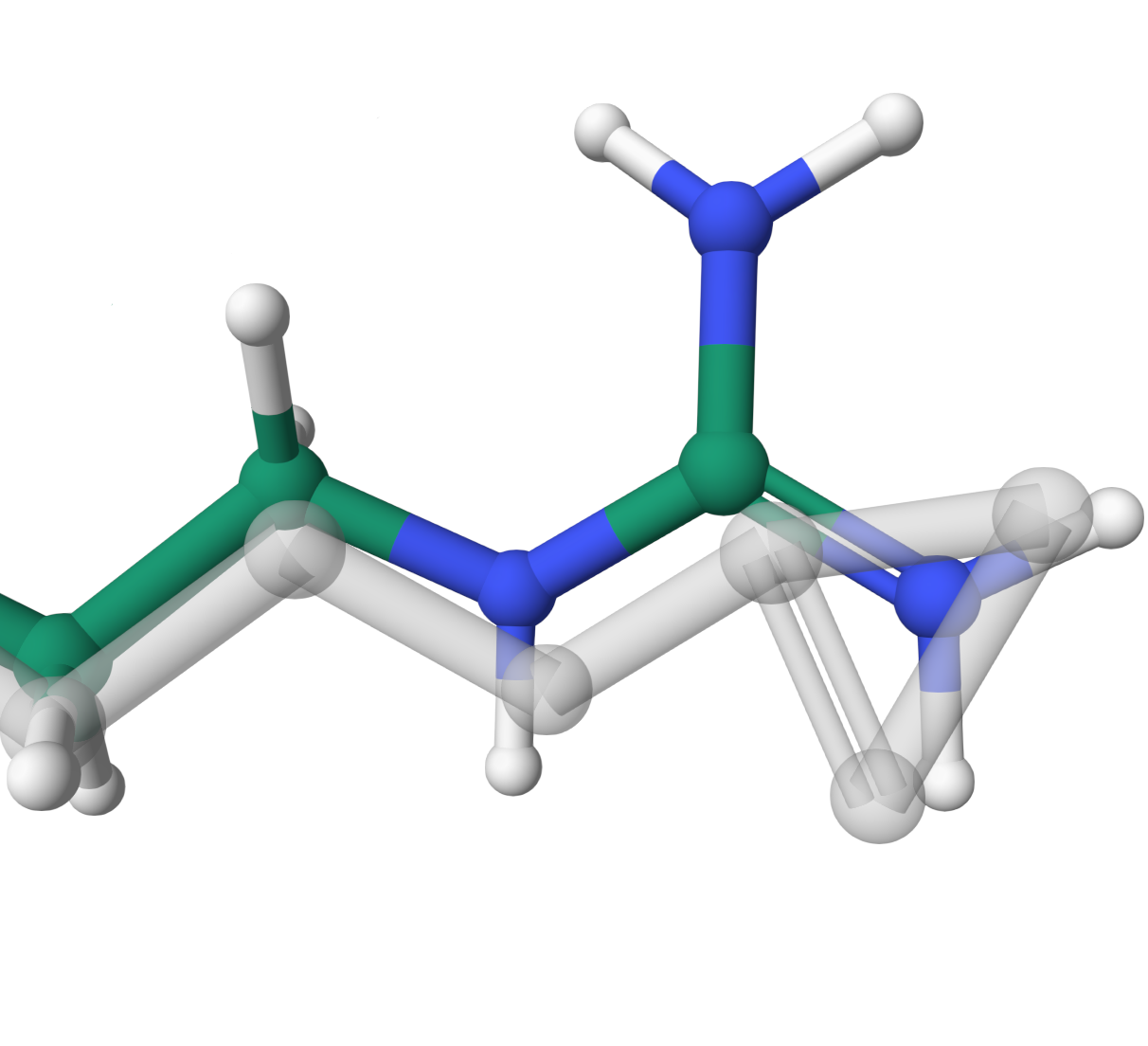
Physically unrealistic
Fixing non-physically predicted atoms NH1 and NH2 of ARG 22 in the structure A4QJE9
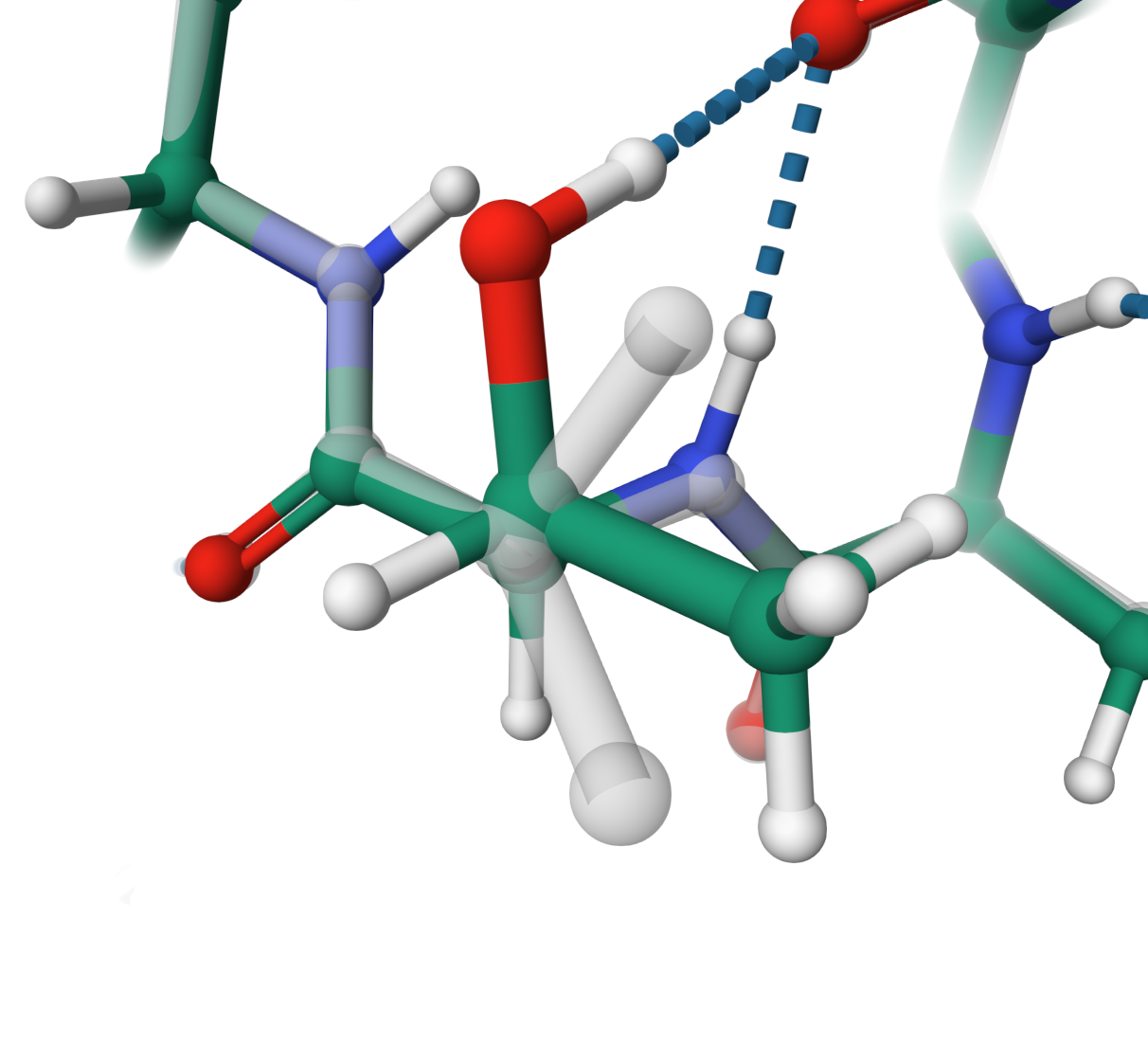
Dihedral angles
Optimisation of THR 555 from an eclipsed conformation in the structure Q57N56
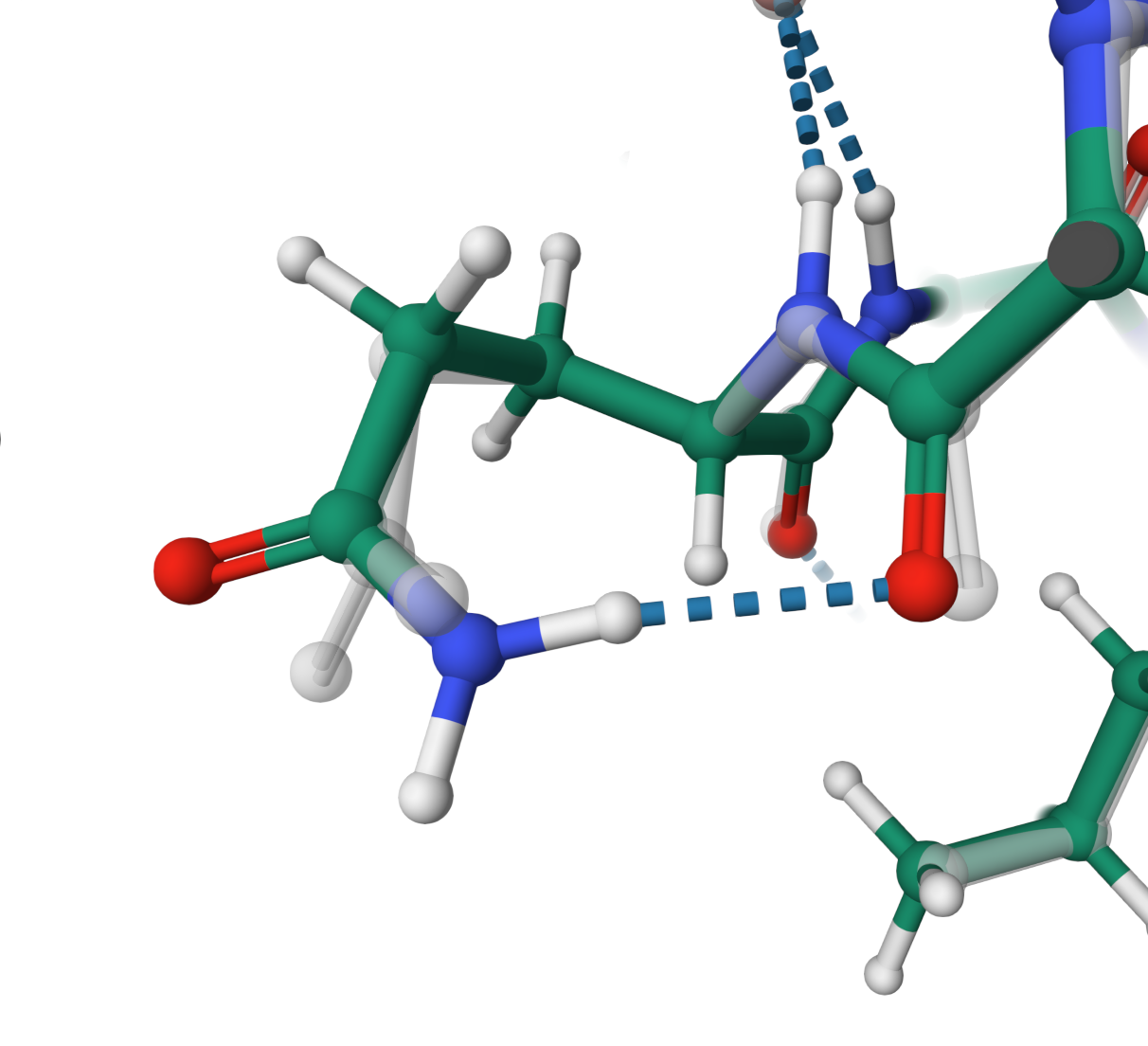
Hydrogen bond
Hydrogen bond formation between ARG 369 and GLN 370 in the structure B7ZW16
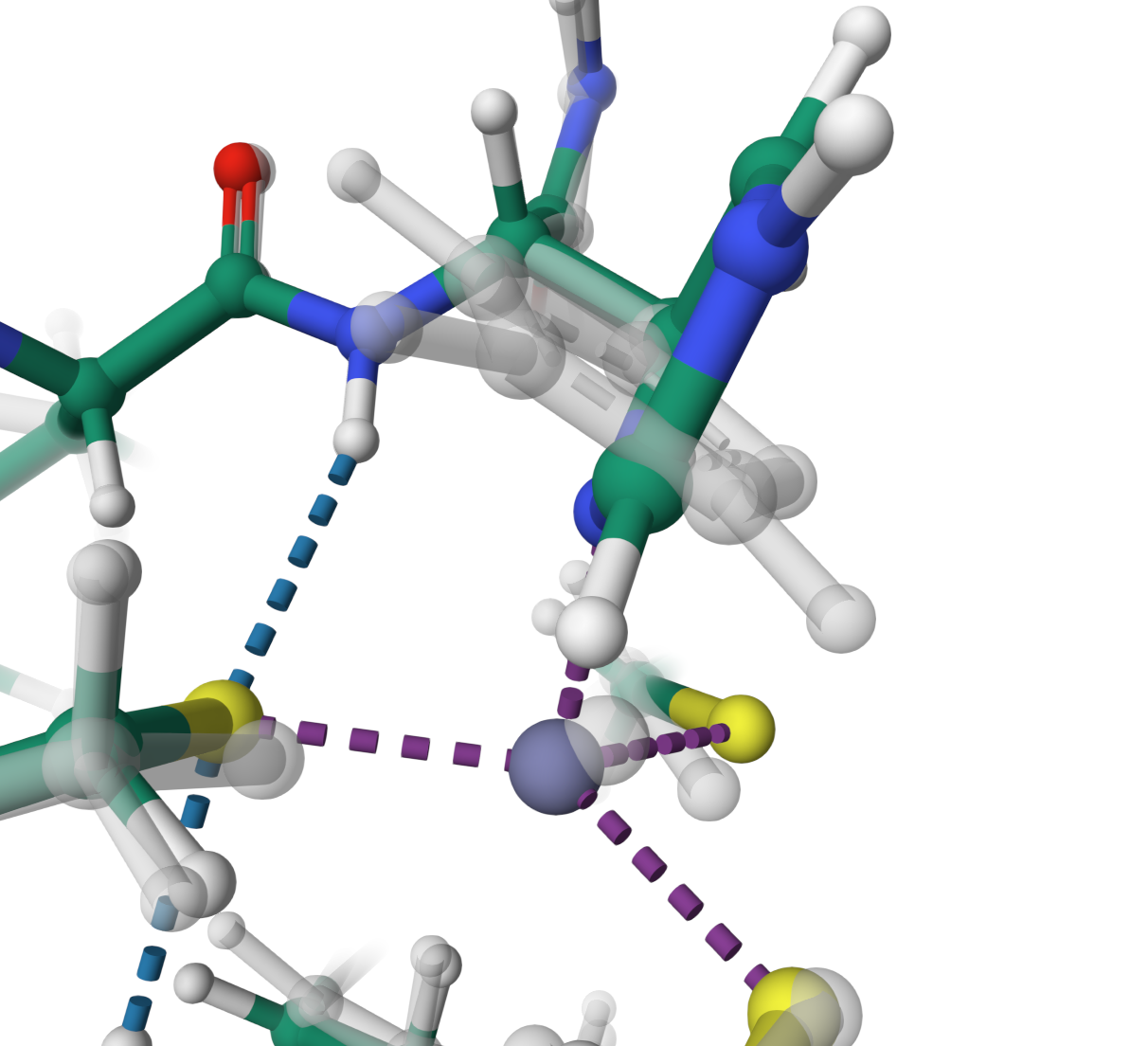
Zinc finger
Optimisation of HIS 25 to achieve a coplanar orientation with the zinc ion in the structure 5xht
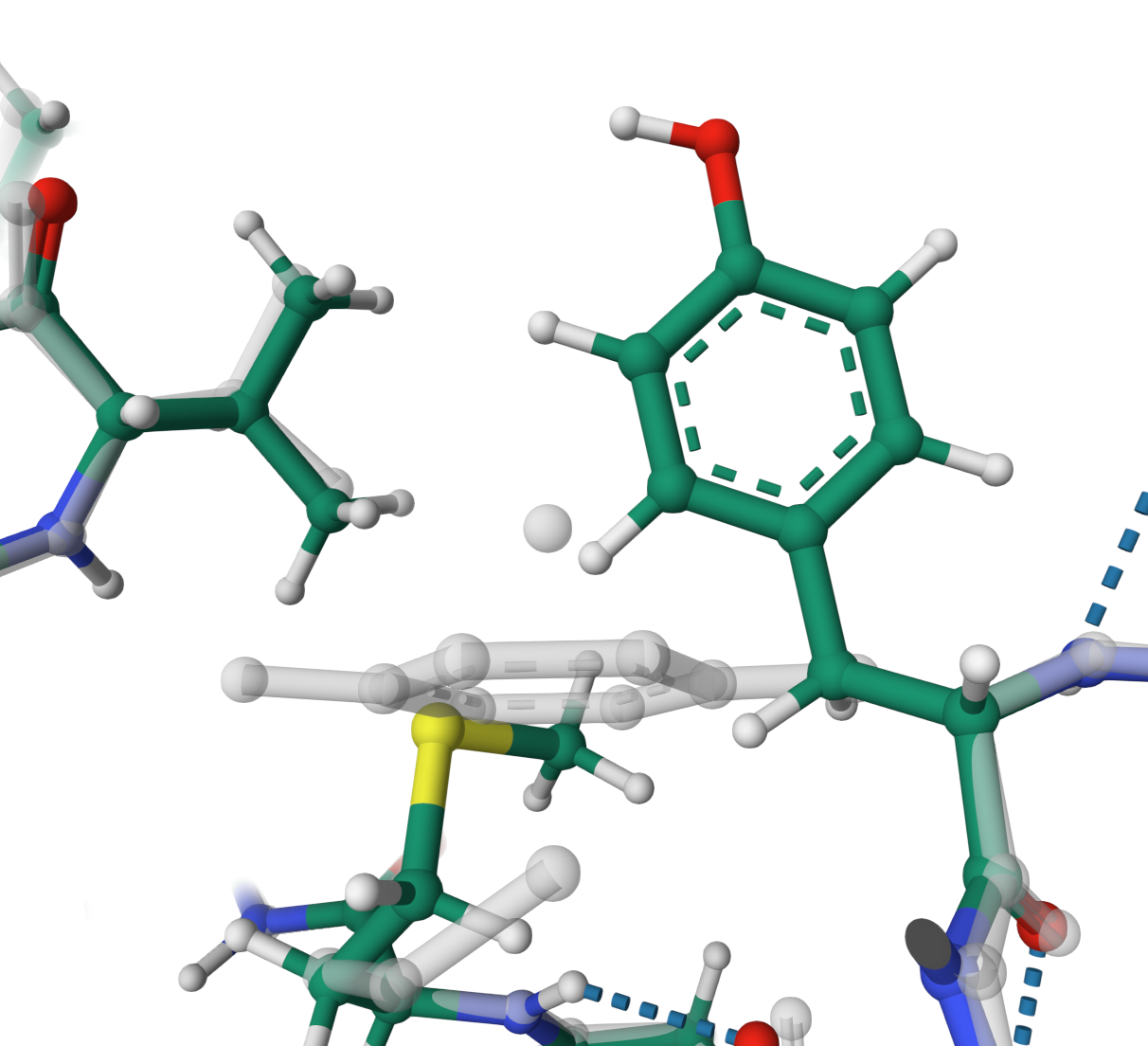
Physically unrealistic
Fixing non-physically predicted atoms of TYR 22 and MET 26 in the structure Q9RS06
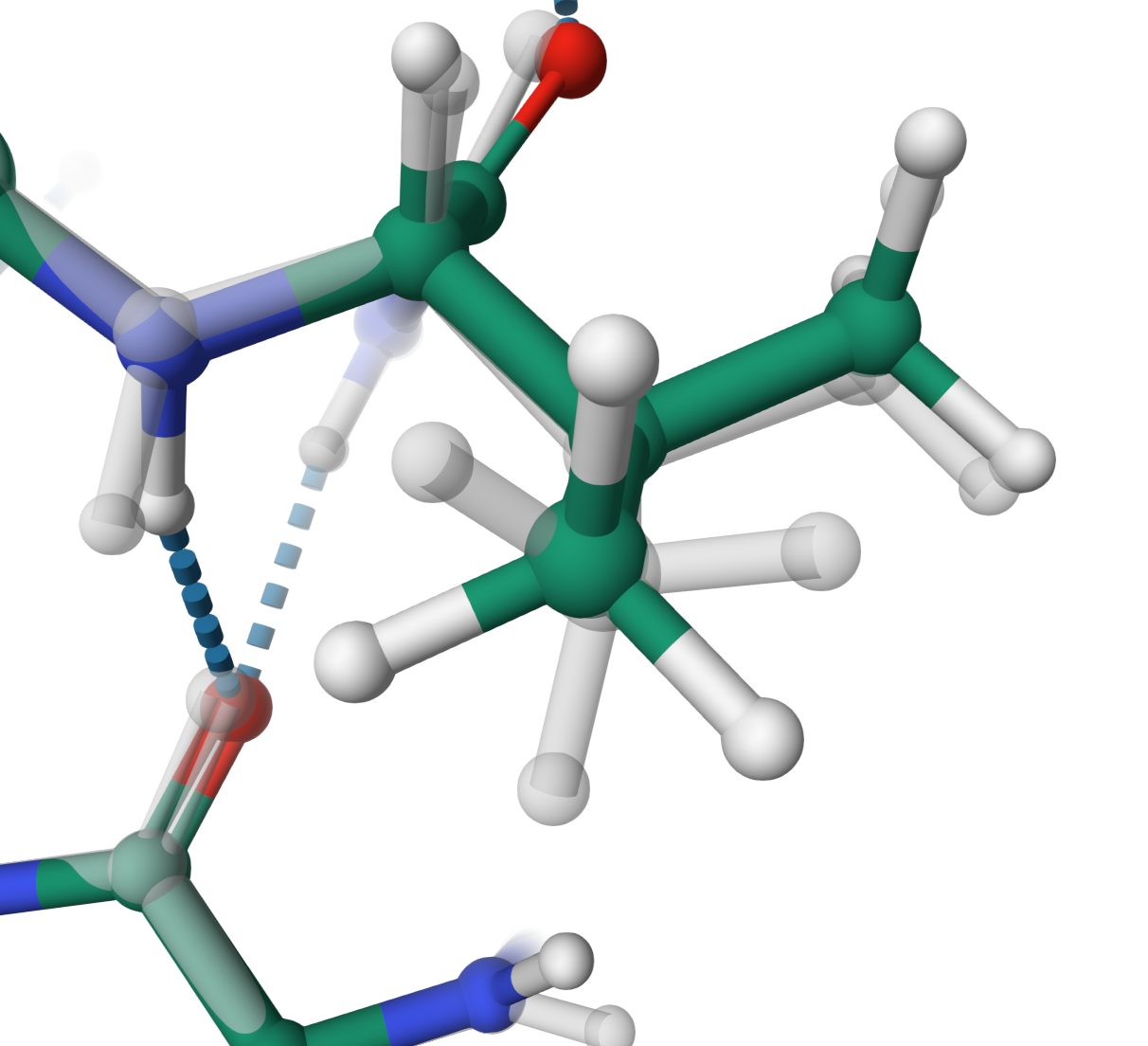
Dihedral angles
Hydrogens optimisation of VAL 123 from an eclipsed conformation in the structure Q3M859
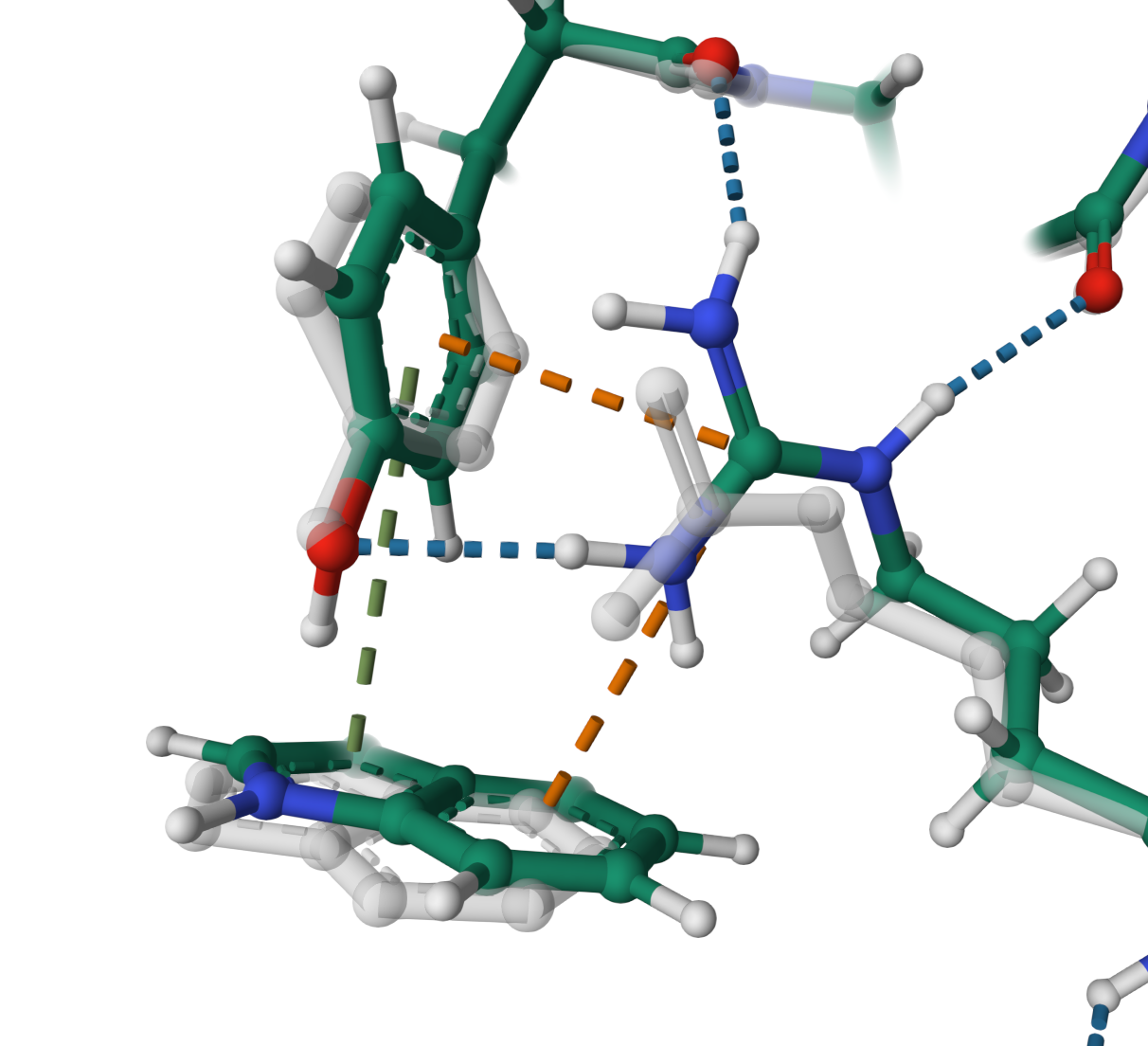
Cation–π interaction
Formation of cation–π interaction of ARG 237 in the structure Q3M859
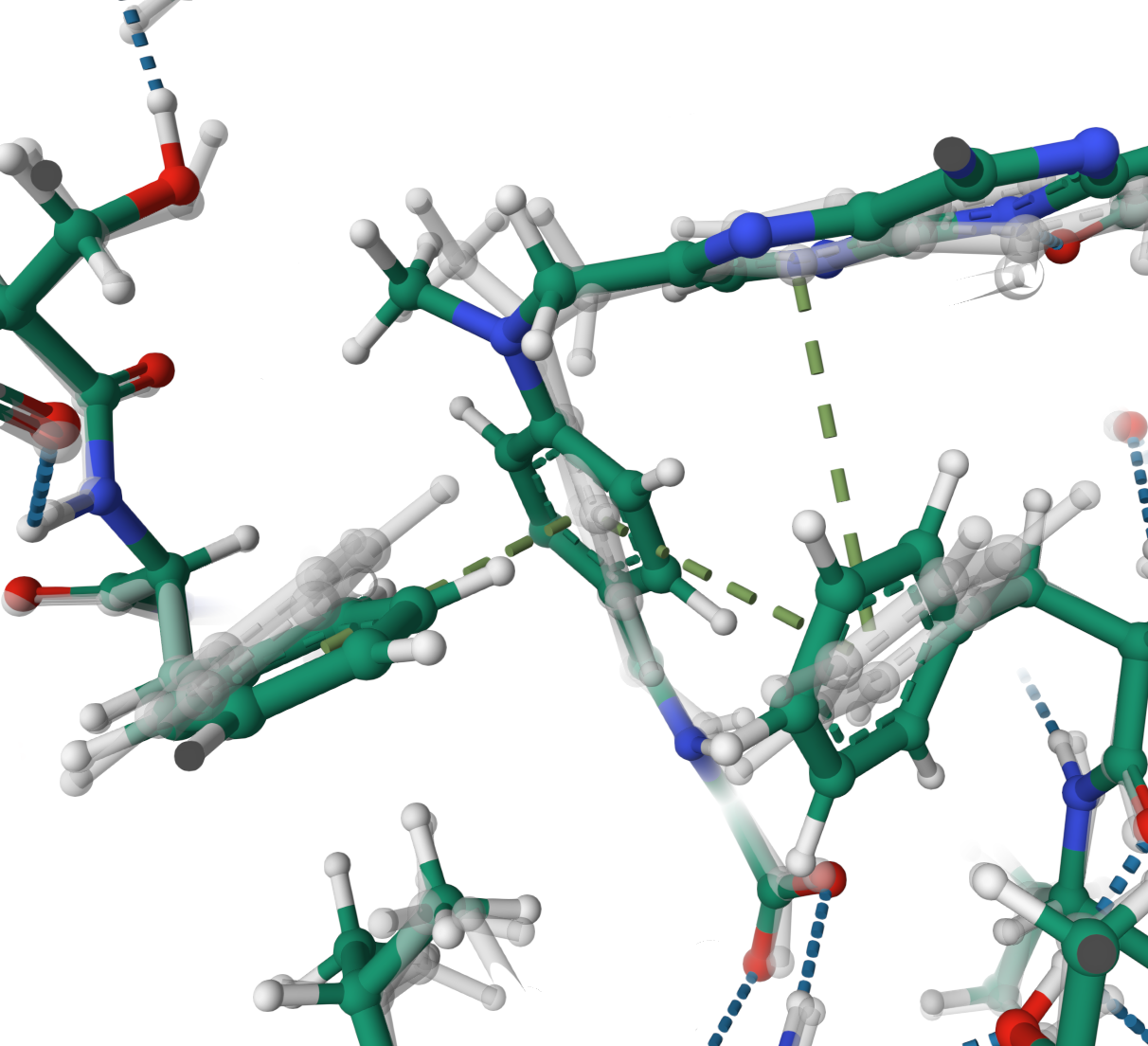
π–π stacking
Stabilisation of MXT 170 via the formation of three π–π stacking interactions in the structure 1ao8
About the RAPHAN approach
Explore our method for fast and accurate protein structure optimisation.

How it work?
Img: Scheme of the RAPHAN approach
PROPTIMUS LIVE optimises protein structures using the RAPHAN method, which is a rapid alternative to optimisation with constrained α-carbons. RAPHAN is an iterative divide-and-conquer method, which divides the protein into overlapping substructures and optimises each substructure separately. Substructures are optimised by physics-based GFN-Force-Field, designed to combine high force-field speed with the accuracy of QM methods.

How accurate is it?
Img: Atom position deviations for RAPHAN and GFN-FF constrained α-carbons optimisations.
The RAPHAN method produces structures comparable to structures optimised by GFN-Force-Field with constrained α-carbons. The mean absolute deviation of atomic positions is approximately 0.03 Å, and the 99th percentile is 0.33 Å. As can be seen in the histogram, structures may differ exceptionally for highly flexible residues.

How fast is it?
Img: Optimisation times for RAPHAN and GFN-FF constrained α-carbons optimisations
A lot. While the duration of GFN-Force-Field optimisation with constrained α-carbons grows approximately quadratically with the size of the structure, by dividing the protein into substructures, RAPHAN's calculation speed is linear with respect to the structure's size. Thanks to parallelisation, PROPTIMUS LIVE optimises proteins without ligands at an average speed of 1000 atoms per minute.
License
Legal Framework
Licence conditions in accordance with § 11 of Act No. 130/2002 Coll. The owner of the software is Masaryk University, a public university, ID: 00216224.
Usage Rights
Masaryk University allows other companies and individuals to use this software free of charge and without territorial restrictions in usual way, that does not depreciate its value. This permission is granted for the duration of property rights.
Terms & Conditions
This software is not subject to special information treatment according to Act No. 412/2005 Coll., as amended. In case that a person who will use the software under this licence offer violates the licence terms, the permission to use the software terminates.